-Search query
-Search result
Showing 1 - 50 of 93 items for (author: krebs & as)

EMDB-16403: 
Subtomogram averaging of a coronavirus spike (ChAdOx1 19E6) in C3 symmetry
Method: subtomogram averaging / : Ni T, Mendonca L, Zhu Y, Zhang P

EMDB-16404: 
Subtomogram averaging of a coronavirus spike (ChAdOx1 19E6) in C1 symmetry
Method: subtomogram averaging / : Ni T, Mendonca L, Zhu Y, Zhang P

EMDB-16405: 
Subtomogram averaging of a coronavirus spike (ChAdOx1 19E) in C3 symmetry
Method: subtomogram averaging / : Ni T, Mendonca L, Zhu Y, Zhang P

EMDB-16406: 
Subtomogram averaging of a coronavirus spike in situ (ChAdOx1 19E) in C1 symmetry
Method: subtomogram averaging / : Ni T, Mendonca L, Zhu Y, Zhang P

EMDB-16697: 
Subtomogram averaging of a coronavirus spike (ChAdOx1 19E6) in C1 symmetry after 3D classification
Method: subtomogram averaging / : Ni T, Mendonca L, Zhu Y, Zhang P

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO

EMDB-16511: 
HERV-K Gag immature lattice
Method: subtomogram averaging / : Krebs AS, Liu HF, Zhou Y, Rey JS, Levintov L, Perilla JR, Bartesaghi A, Zhang P

EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
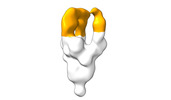
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
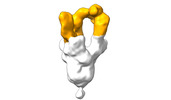
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
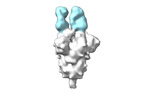
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
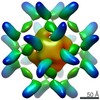
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-13292: 
Cryo-EM structure of the MoStoNano fusion protein
Method: single particle / : Benoit RM, Poghosyan E

EMDB-24346: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-80 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24348: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-081 Fab
Method: single particle / : Saphire EO, Li H

EMDB-24349: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-091 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24350: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-094 VHH-Fc
Method: single particle / : Saphire EO, Li H

EMDB-24351: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-096 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24358: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-147 IgG
Method: single particle / : Saphire EO, Yu X

EMDB-24359: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-246 IgG
Method: single particle / : Saphire EO, Rayaprolu V

EMDB-24335: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-259 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24336: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-186 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24337: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-199 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24338: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-249 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24339: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-252 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24340: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-049 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24341: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-073 scFv
Method: single particle / : Saphire EO, Li H

EMDB-24342: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-043 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24343: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-010 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24344: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-148 IgG
Method: single particle / : Saphire EO, Li H

EMDB-24345: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-002 Fab
Method: single particle / : Saphire EO, Li H

EMDB-24352: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-250 Fab
Method: single particle / : Saphire EO, Li H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
